C++ Shortest Path - Dijkstra PQ Implementation, Bellman-Ford Code As Backup, and Floyd Warshall4/5/2018
Introduction: Shortest Paths
We've already had a blog post on graph theory, adjacency lists/matrixes for representing a graph, and BFS/DFS for traversing a graph. So how do we find the shortest path in a graph if there are edge weights? The answer is...we use the famous Dijkstra algorithm. (Assuming there are no negative edge weights.) If there are negative edge weights, then we use Bellman Ford.
What we want is, for a starting vertex/node, a list containing the shortest path from the starting node to every other node.
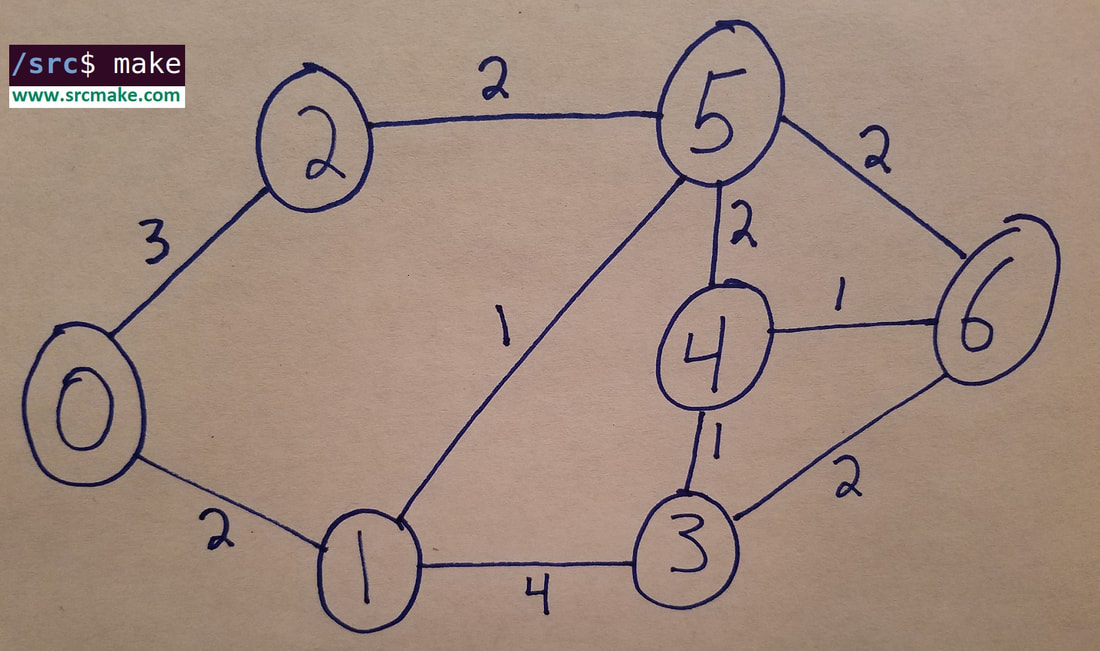
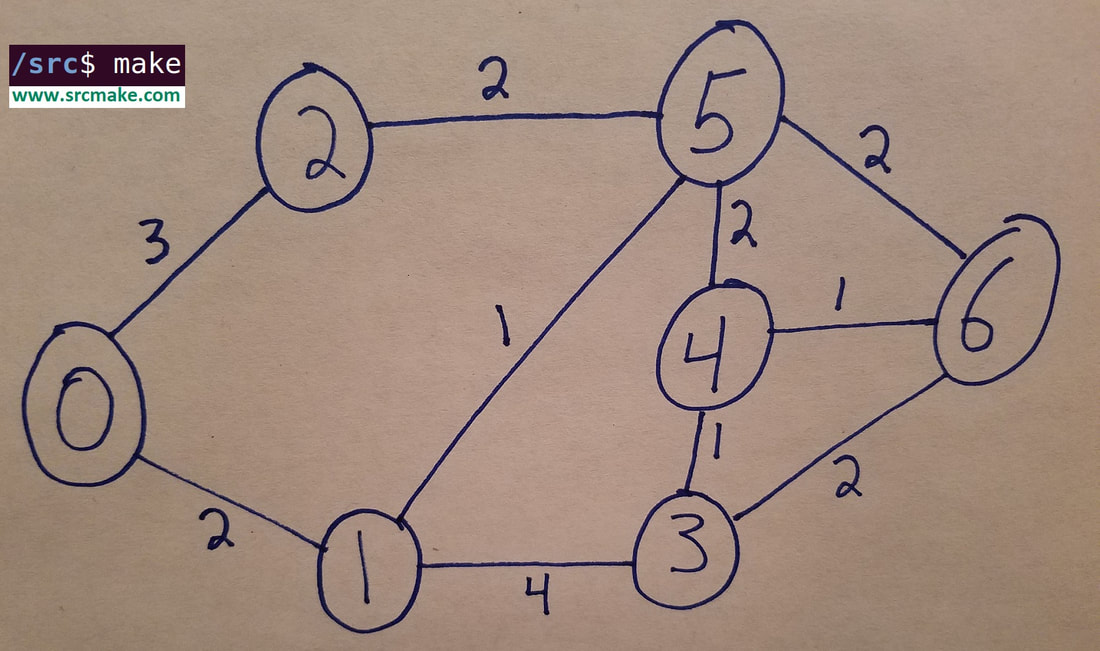
The above graph is what we'll be using in our algorithms. You can see that there are some nodes (vertices) labeled 0 to 6, and there are some edges that each have weights on them. We want the shortest path (lowest weight total) from one node to another. To the right is the adjacency list representation of the graph, where for adjList[i], there's a vector of pair<int, int>s at each index, where the first int represents the neighbor node, and the second int represents the edge weight between them.
Djikstra's Algorithm For Shortest Paths, using Priority Queue
Djikstra's algorithm is going to maintain a list of shortest paths from the starting node to all other nodes. Initially, the starting node has 0 distance to itself, but infinity distance to all other nodes. Then, by looking at all neighbors, of the node we're processing (starting with the start node), we'll update the shortest distances by checking if we can do better.
In this implementation, we use a priority queue, which will prioritize checking the shortest edge weights first.
Our code will generate that adjacency list, and perform the algorithm we mentioned before.
Bellman-Ford - Good For Negative Edges
Dijkstra is a greedy algorithm and will fail if there are cycles or negative edge weights. Because of this, we need the Bellman-Ford algorithm in that case. (But it's slower than Dijkstra, so use Dijkstra depending on the problem constraints.)
The results of running this should be exactly the same as Dijkstra. (But this takes longer.)
Floyd Warshall - Efficient For All Sources To All Nodes
Dijkstra and Bellman-Ford calculate the shortest path in a graph from one source node to all other nodes. But what if we need to calculate the shortest distance from all sources as the starting node, to all other nodes? That's what the Floyd Warshall algorithm is for.
We'll need an adjacency matrix representation of our graph for this problem. This time, edge weights that don't exist are set to infinity, and the distance of a node from itself is 0.
The output for node 0 is the same as Dijkstra and Bellman-Ford.
To Get The Path Itself
In our code, we saved the distance for each node from the source. But what if we need to know the actual shortest path (meaning what the nodes are, and not just the distance)?
Basically, in our distance list, we store not only the distance from "source node" to "current node", we also store the "previous node" in an int variable. Here's some updated code to show Dijkstra's algorithm where paths are stored.
The changes are to "vector<int> dist"" to "vector< pair<int, int> > dist" in our Dijkstra function (as well as line 116 where we store the previous node), the main function where the result is returned, and in the print function.
Conclusion
It takes some time to understand shortest path algorithms, but with some practice, it becomes second nature.
Next up in the graphing algorithm series is the Ford-Fulkerson algorithm for Maximum Flow. Check out this bloc post to see it.
Like this content and want more? Feel free to look around and find another blog post that interests you. You can also contact me through one of the various social media channels.
Twitter: @srcmake Discord: srcmake#3644 Youtube: srcmake Twitch: www.twitch.tv/srcmake Github: srcmake Comments are closed.
|
AuthorHi, I'm srcmake. I play video games and develop software. Pro-tip: Click the "DIRECTORY" button in the menu to find a list of blog posts.

License: All code and instructions are provided under the MIT License.
|